ESciDoc Services Annotator
Introduction[edit]
The motivation for defining an Annotator service comes from MPI MB Demonstrator proposal. Main aim is to be able to link publications with references or short descriptions/comments on resources about which a publication (or another form of work) is about. There is an assumption that most of these are external resources and are not managed by MPDL services. However, this does not have to be the case.
Use cases[edit]
- MPI Marine Microbiology use cases
Related concepts/implementations[edit]
Basic Data Model[edit]
- For basic data model and definitions please check Open Annotation Collaboration (OAC)
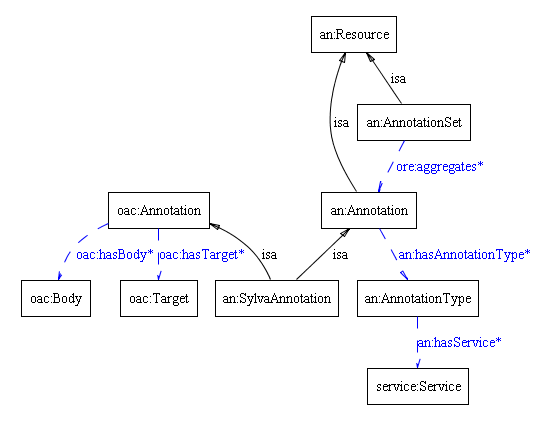
- The ontology developed for the Annotator service at MPDl is shown below
- A Resource is a class of all individuals representing content which is managed in the system. In the depicted ontology, Annotation(SilvaAnnotation) and Target are considered as a Resource.
- An Annotation is a subclass of managed Resource class. It is class of individuals that relate the Body and the Target individuals, and may have some additional properties.
- A Body is the class of individuals which are considered as resources that annotate the Target individual.
- A Target is the class of individuals which are annotated.
- A SilvaAnnotation is a subclass of an Annotation referencing Silva database resources (see MPDL Demonstrator Marine Microbiology for more details)
- A Service is a class of individual services that may be associated with particular types of annotations, e.g. Silva browser, GeoBLAST
- Notes
- In particular cases an Annotation may not have related Body individual. In this case for the Annotation individual it is assumed that it is an e.g. "textual" annotation (e.g. it is described via a "datatype" property as a content of the annotation rather than an "object" property).
- this model makes use of the OAC Baseline model. As at present more details on the model are not clear, additional Annotation class is defined in the MPDL Annotation ontology. The SilvaAnnotation is thus a subclass of both an:Annotation and oac:Annotation classes. The an:Annotation class in addition models relations to the Annotation types and services related for particular annotation type. (The model will be aligned with the OAC Baseline model as soon as more details are known).
Data Example[edit]
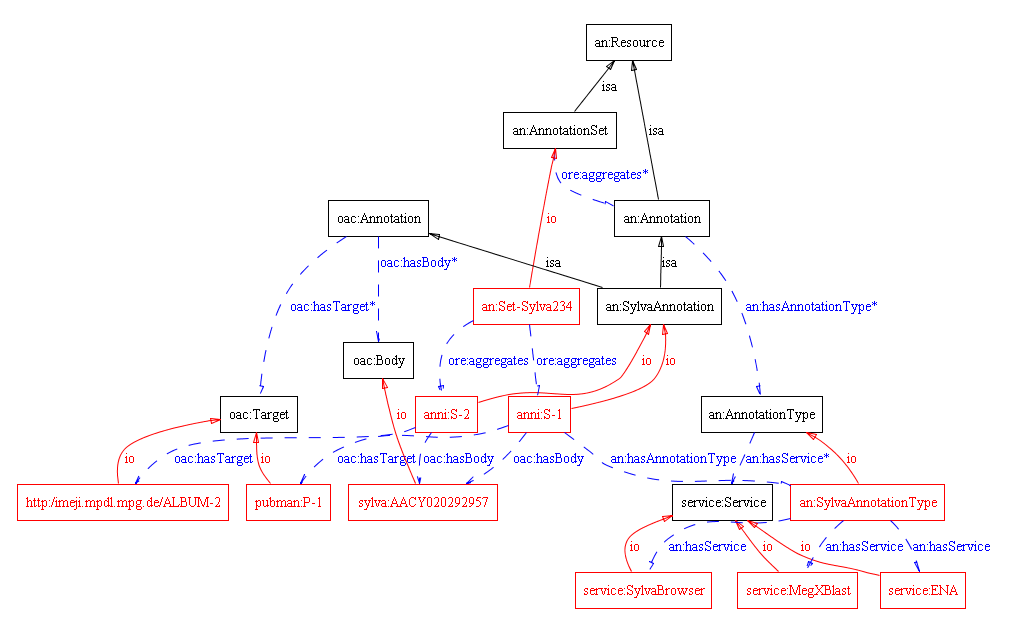
- pubman:P-1 is a publication about a "Marine metagenome 1096626369196, whole genome shotgun sequence".
Environmental Genome Shotgun Sequencing of the Sargasso Sea,
J. Craig Venter,Karin Remington,John F. Heidelberg,Aaron L. Halpern,Doug Rusch,Jonathan A. Eisen,Dongying Wu,Ian Paulsen,Karen E. Nelson,William Nelson,
Derrick E. Fouts,Samuel Levy,Anthony H. Knap,Michael W. Lomas, Ken Nealson, Owen White,Jeremy Peterson,Jeff Hoffman,Rachel Parsons,Holly Baden-Tillson,
Cynthia Pfannkoch, Yu-Hui Rogers, and Hamilton O. Smith
Science 2 April 2004: 66-74.Published online 4 March 2004, http://www.sciencemag.org/content/304/5667/66
- silva:AACY020292957 is the resource in the Silva database, providing some more details on the sequence (see http://www.arb-silva.de/browser//AACY020292957 ). A Target individual may have additional properties. These are highly dependant on the target itself.
- anni:S-1 is a SilvaAnnotation that has as a body the pubman:P-1 publication in PubMan repository and as a target the silva:AACY020292957 resource in Silva database. This individual has properties refering to the environmental data such as geo coordinates, some description and comment, system generated properties such as: created-by, created-on.
- an:SilvaAnnotationType is an individual of the class AnnotationType and represents an instance of Annotation types related to Silva resources.
- service:MegXBlast and service:SilvaBrowser are concrete services of which the system is aware of. As being related to the an:SilvaAnnotationType via an:hasService property, it is clear that these are services that can derive some functionality for the annotations which are related to an:SilvaAnnotationType individual. For example: MegXBlast algorithm, transformation and automated population of some annotation properties via SilvaBrowser service etc. (Note: the example assumes that the SilvaBrowser may offer data in processable format, to be checked with the institute directly - here provided only as an example).
Remark[edit]
- To be considered during implementation: check if it makes sense to keep some automatically ingested Target instance properties with the Annotation instance itself or with the Target instance. This will not conflict the basic ontology model, but would probably require subclassig the Target in similar manner as the Annotations.
Data model extensions[edit]
Annotation set[edit]
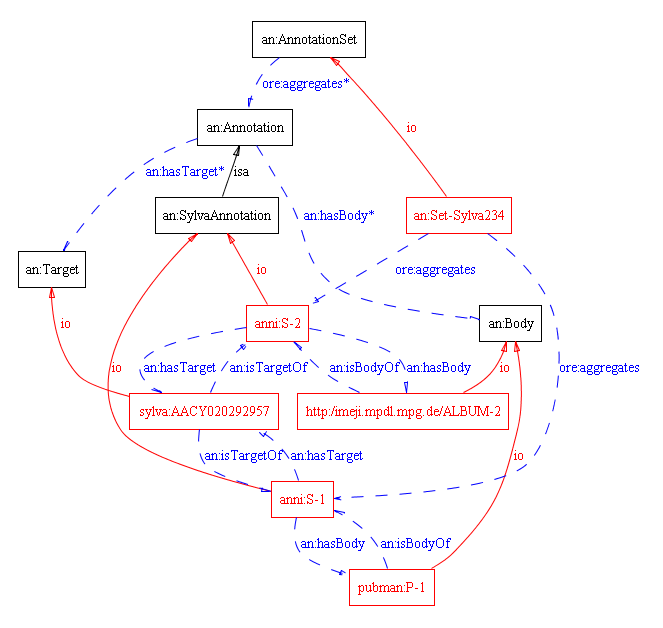
In particular cases there is a need to related various Annotations into an Annotation set. An Annotation set can be considered is an arbitrary choice of Annotations related to a particular topic, created by a particular user or user group (in case of Collaborative annotations). An extension to the model that includes also Annotation sets is given below.
For the purpose of relating the Annotations into an AnnotationSet, OAI-ORE term "aggregates" is used (see OAI-ORE Vocabulary )
Implementation concept[edit]
The Annotator service implementation has two general components:
- Annotator store
- Annotator tool
Note: concrete implementation may bring other aspects on this issue, therefore the concept provided here shall be considered as a high-level design.
Annotator store[edit]
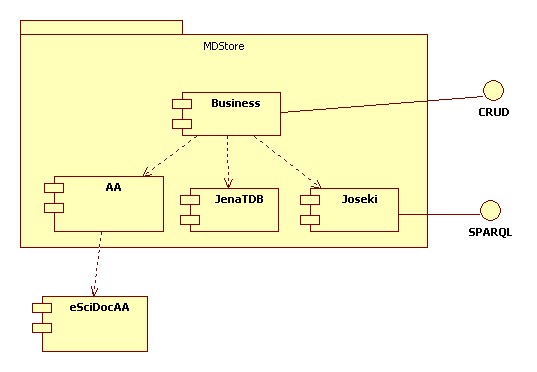
- The Annotator store is an RDF store offering interfaces to create and query the data. An architecture similar to the MDStore Architecture is envisioned for this purpose.
- The RDF Store is primary storage for the Annotator resources
- eSciDoc core infrastructure and FedoraCommons repository may be used as an LTA archival solution - however, this would be completely separated from the Annotator store.
- Internal AA component will be built in the Annotator store, however pluggable to implement use external AA components such as eSciDoc AA.
Annotator tool[edit]
- The Annotator tool is the user interface that helps creating various types of annotations.
As there are plenty of possibilities on how an annotation may be created (depending on the type of resources, particular user scenario etc.) it is very heavy to decide upon a single annotation tool. It is a matter of choice of the end user.
- As the Annotator store interfaces and data model is based on the OAC data model, in general, any tool that would support the OAC data model may be used as an Annotator tool, with small or no modification needed. (keeping in mind that the Annotator store is exposed as a REST service interface and a SPARQL end point - practically a tool which e.g. creates serialized OAC RDF/XML or implementing the SPARQL interface properly can be used to write to the Annotator store).
Annotator store and Linked data publishing[edit]
- The MD Store and the Annotator store implementations may be used as tools to have published Linked data on e.g. content resources maintained in eSciDoc such as : publications, images, manuscripts, annotations etc.
- Providing proper searching and querying interfaces to linked data brings other possibilities and allows finding new facts out of the existing data - to be used for cross disciplinary analysis
- This case needs to be carefully analyzed form the aspect of scalability and performance of available triple store and linked data platforms (see Triplestores, Linked data tools)
- As potential starting point, selected data collections may be used to showcase the implementation